Gene annotation of the human UROD gene
Practical exercise
Enrique Blanco - eblanco@imim.es
Roderic Guigo - rguigo@imim.es
|
Abstract: In this exercise, a previously annotated gene will be used to
measure the accuracy of different gene finding approaches. geneid,
GENSCAN, FGENESH and GENEWISE will be used to annotate the
sequence. Both search by signal, content and homology (protein and cDNA sequences)
methods will be employed in order to improve the ab initio results. Weak conservation
in Start codons and the short length of the initial exon leads to wrong prediction of
this exon in most cases.
Colour legend:
Genomic element
Operations or links
|
|
Gene prediction (T)
|
Gene structure in Eukaryotes:
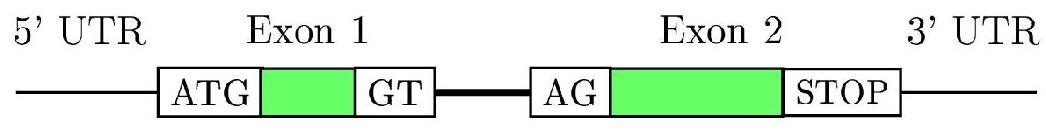
Signals and regions:
| Signals |
| Start (ATG) |
| STOPs (TGA,TAA,TAG) |
| Donor (GT) |
| Acceptor (AG) |
|
|
| Exons |
| Single |
| First |
| Internal |
| Terminal |
|
|
| Regions |
| Exons |
| Introns |
| Intergenic |
| 5' and 3' UTRs |
|
Predicted exons:
geneid structure:
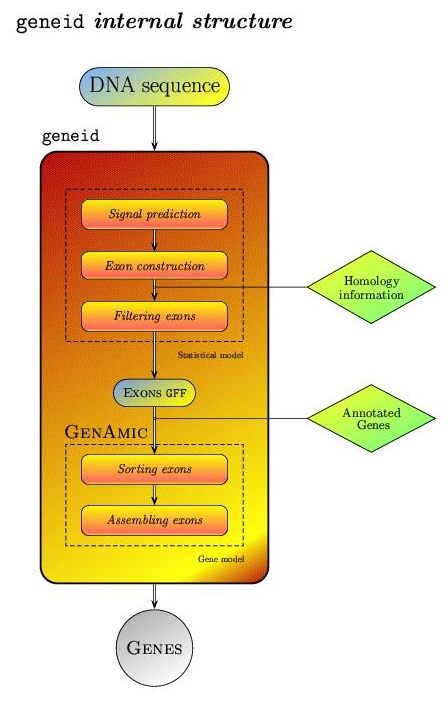
|
|
A. Gene annotation
|
Step 1. Accesing EMBL database to retrieve the gene
- Go to EMBL database
- Select Get Nucleotide sequences
- Type sequence name U30787
- Press Go button
- Click on the EMBL:U30787 (coding sequences) entry link
- Have a look at the different entry fields: detect the mRNA and CDS exons
- Select Text Entry to see the plain text file
- Get back to the query results
- Click on the EMBL:U30787 (Primary Accession) entry link
- Look at the archive of versions of this entry.
|
|
B. Exploring ab initio gene prediction
|
Step 2. Running geneid
- Connect to the geneid server
- Paste the FASTA sequence
- Choose geneid output format
- Run geneid in diffent levels (compare with the EMBL annotations):
- Searching signals: Select acceptors, donors, start and stop codons. Look for them in the real annotation of the sequence
- Searching exons: Select All exons and try to find the real ones
- Finding genes: You do not need to select any option (default behaviour). Compare the predicted gene with the real gene
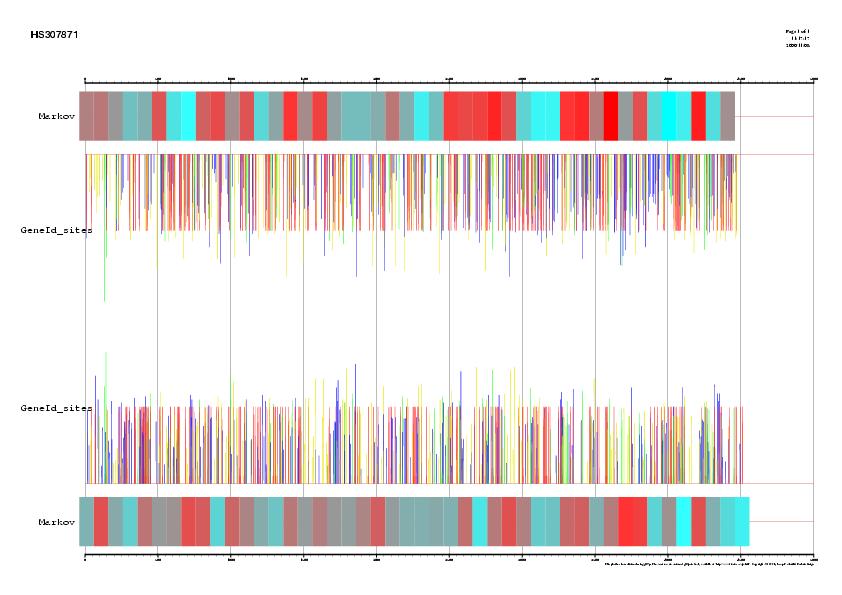 |
 |
 |
|
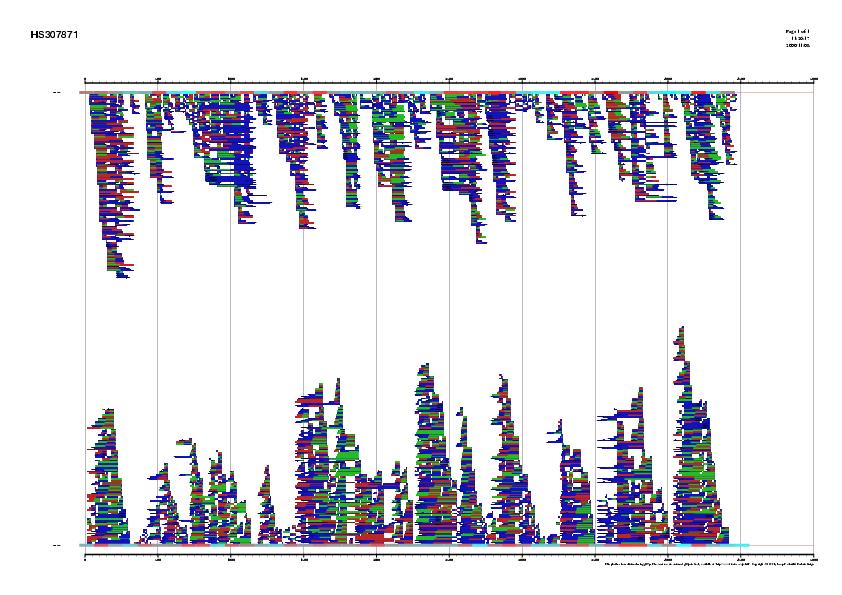
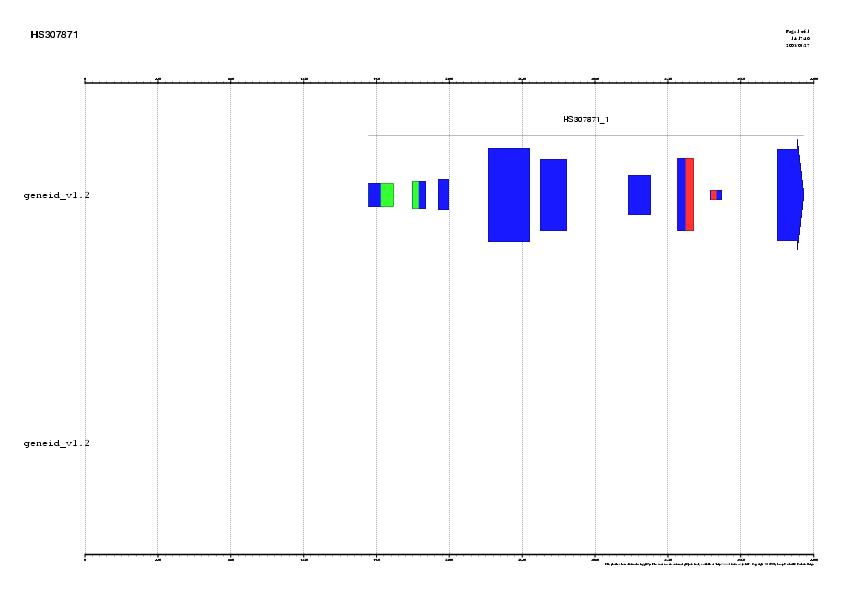
Figure 1. Signal, exons and genes predicted by geneid in the
sequence HS307871
|
- Try to get the gene prediction again, but using the parameter file for:
Drosophila melanogaster, Caenorhabditis elegans and Oryza sativa.
Can you see something in common? Do you realize species-specific training is not so
specific?
|
Step 3. Running other genefinders
As there are several alternative programs to analyze a DNA
sequence, we can run each application and observe the common parts of the
predictions.
- GENSCAN:
- Connect to the GENSCAN server
- Paste DNA sequence
- Press Run Genscan button
- Compare annotations and predictions
- FGENESH:
- Connect to Softberry homepage
- On the left frame, select GENE FINDING in Eukaryota
- Select the program FGENESH
- Paste DNA sequence
- Press Search button
- Compare annotations and predictions
- NOTE: First exon is always missed in the predictions and there are some
problems to detect the donor site from exon 5. Detection of Start codons is
a serious drawback in current gene finding programs (see Figure 2). However,
this problem can be overcome by using homology information to complete the
gene prediction.
|
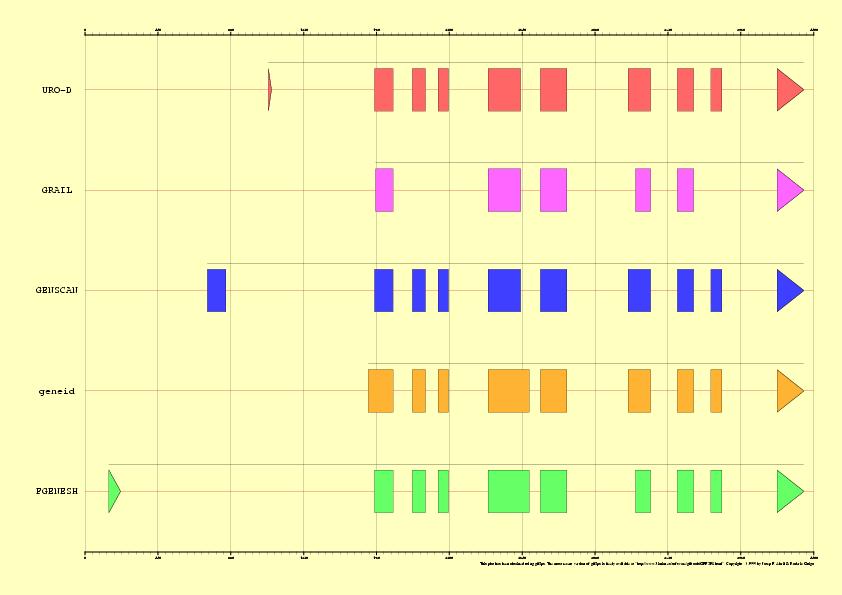
Figure 2. EMBL annotation and genes predicted by Grail,
GENSCAN, geneid and FGENESH in the sequence HS307871
|
|
|
C. Using EST/cDNA homology information
|
Step 4. Using FGENESH-C + alignment of transcripts
Let's search for ESTs supporting future predictions by FGENESH-C:
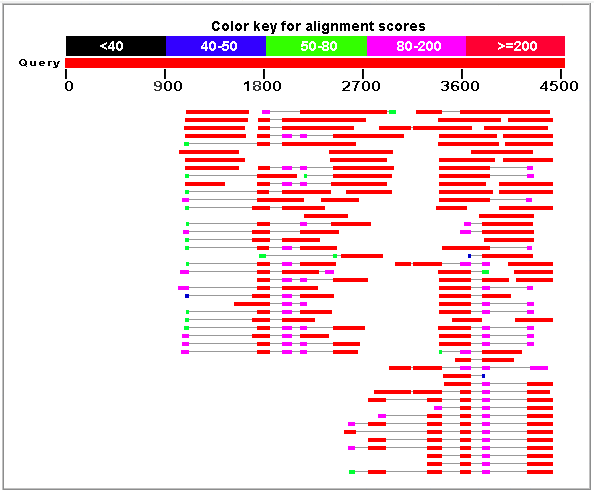 |
|
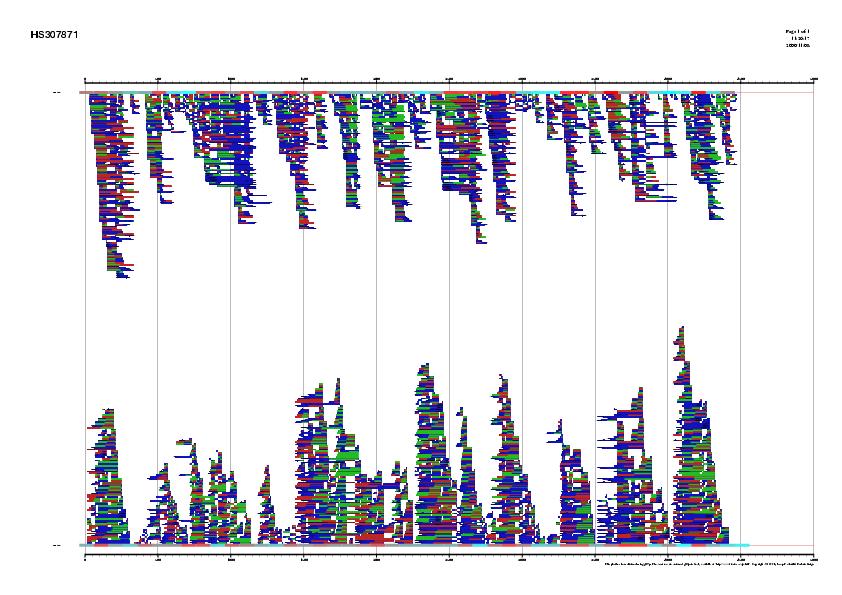
Figure 3. Best human ESTs in the alignment mapped on the genomic sequence HS307871
|
- Connect to the FGENESH-C server
(on Gene finding with similarity menu)
- Paste the sequence U30787
- Paste the DA976126 SYNOV2 EST (the last one in the first column)
- Press the Search button
- Analyze the prediction (compare to the annotations)
|
|
D. Using protein homology information
|
Step 5. Spliced alignment
Spliced alignment is very useful when we have additional information
(a putative homologous protein sequence) about the content of the sequence.
Thus, gene prediction is guided by fitting the protein sequence into the
best splice sites predicted in the genomic sequence.
- Open the NCBI blast server
- Choose blastx program
(genomic query versus protein database)
- Paste the genomic sequence and press the
Blast! and
Format!
- Select the first protein. Go to the GenBank entry and convert it into a
FASTA sequence (it is the real protein annotated in the genomic sequence).
- Open genewise web server
to use this protein to predict the best gene structure
- Paste both protein and genomic sequences and run the program
- Compare predicted gene (end of the file) and annotations: look
for splice sites within introns to check exon boundaries are correct
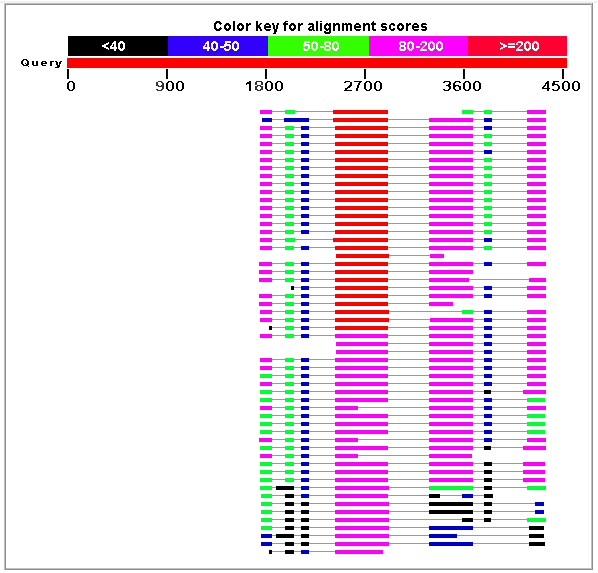 |
Figure 5. Best HSPs representing proteins homologues similar to the
genomic sequence HS307871 obtained using blastx
|
|
|
|
E. Results
|
Here you can find the solutions to every exercise:
|
|
|
F. Bibliography
|
- J.F. Abril and R. Guigó.
gff2ps: visualizing genomic annotations. Bioinformatics 16:743-744 (2000).
- Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215:403-410 (1990).
- Burge, C. and Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78-94 (1997).
- E. Blanco, G. Parra and R. Guigó.
Using geneid to Identify Genes. In A. D. Baxevanis and D. B. Davison,
chief editors: Current Protocols in Bioinformatics. Volume 1, Unit 4.3.
John Wiley & Sons Inc., New York. ISBN: 0-471-25093-7 (2002).
- Asaf A. Salamov and Victor V. Solovyev. Ab initio Gene Finding in
Drosophila Genomic DNA Genome Res. 10: 516-522 (2000).
-
D. Hyatt, J. Snoddy, D. Schmoyer, G. Chen, K. Fischer, M. Parang, I. Vokler, S. Petrov, P. Locascio, V. Olman, Miriam Land, M. Shah, and E. Uberbacher.
Improved Analysis and Annotation Tools for Whole-Genome Computational Annotation and Analysis: GRAIL-EXP Genome Analysis Toolkit and Related Analysis Tools.
Genome Sequencing & Biology Meeting (2000).
- Ewan Birney and Richard Durbin. Using GeneWise in the Drosophila
Annotation Experiment. Genome Res. 10: 547-548 (2000).
|








