Sequences : PWM : JPG / PNG / PS
Sequences : PWM : JPG / PNG / PS

Sequences : PWM : JPG / PNG / PS
Sequences : PWM : JPG / PNG / PS
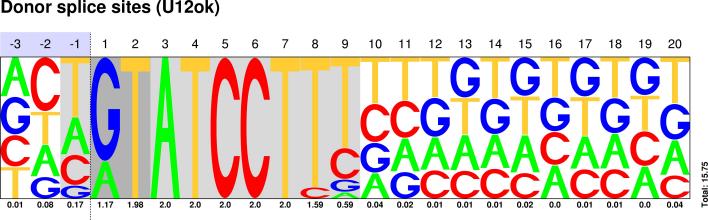
Sequences : PWM : JPG / PNG / PS
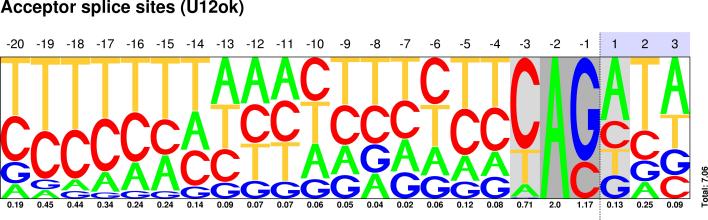
Sequences : PWM : JPG / PNG / PS
POINT
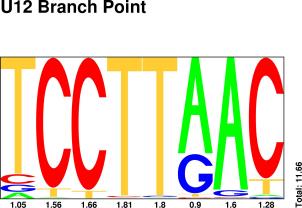
Sequences : PWM : JPG / PNG / PS
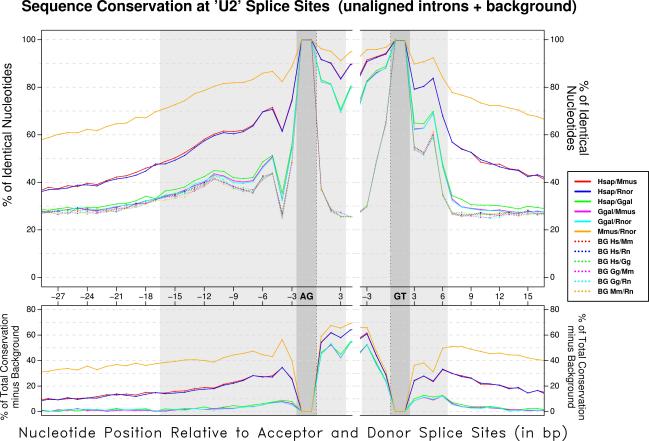
We have carried out an initial analysis of the dynamics of the recent evolution of the splice sites sequences on a large collection of human, rodent (mouse and rat), and chicken introns. Our results indicate that the sequences of splice sites are largely homogeneous within tetrapoda. We have also found that orthologous splice signals between human and rodents and within rodents are more conserved than unrelated splice sites, but the additional conservation can be explained mostly by background intron conservation. In contrast, additional conservation over background is detectable in orthologous mammalian and chicken splice sites. Our results also indicate that the U2 and U12 intron classes seem to have evolved independently since the split of mammals and birds; we have not been able to find a convincing case of interconversion between these two classes in our collections of orthologous introns. Similarly, we have not found a single case of switching between AT-AC and GT-AG subtypes within U12 introns, suggesting that this event has been a rare occurrence in recent evolutionary times. Switching between GT-AG and the non-canonical GC-AG U2 subtypes, on the contrary, does not appear to be unusual; in particular, T to C mutations appear to be relatively well tolerated in GT-AG introns with very strong donor sites.
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Hsap | UCSC_200307 | 21744 | 20894 | 18117 | 15159 | 10757 | 7799 | 17939 | 15066 | 10316 | 7443 | 21091 |
| Mmus | UCSC_200310mm | 17988 | 16126 | 14432 | 13677 | 9765 | 9010 | 14175 | 13461 | 9078 | 8364 | 16192 |
| Rnor | UCSC_200306rn | 4798 | 4134 | 3454 | 3347 | 2201 | 2094 | 3368 | 3275 | 1947 | 1854 | 4536 |
| Ggal | UCSC_200402 | 1496 | 1085 | - | - | - | - | - | - | - | - | 1367 |
| Hsap | UCSC_20030410 | 19174 | 18337 | 18145 | 18067 | 10486 | 10408 | 18014 | 17901 | 9988 | 9875 | 18226 |
| Mmus | UCSC_200302mm | 13406 | 11161 | 10503 | 10404 | 7397 | 7298 | 10371 | 10255 | 6908 | 6792 | 12511 |
| Rnor | UCSC_200301rn | 4219 | 3372 | 3070 | 3049 | 2102 | 2081 | 3017 | 2991 | 1893 | 1867 | 4002 |
| Based on | All Exons | All Introns | All CDSs | Splice Sites | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| refgenes.txt | SEQ(fasta) | SEQ(content) | SEQ(fasta) | SEQ(content) | SEQ(fasta) | SEQ(content) | EXONIC | INTRONIC | |||
| Hsap UCSC200307 | 19M | 3.7M | 362M | 4.9M | 11M | 4.8M | 19M | 17M | |||
| Mmus UCSC200310 | 15M | 2.8M | 211M | 3.6M | 8.5M | 3.7M | 15M | 14M | |||
| Rnor UCSC200306 | 4.0M | 878K | 70M | 1.1M | 2.6M | 1.1M | 4.7M | 4.4M | |||
| Ggal UCSC200402 | 1.2M | 260K | 13M | 325K | 772K | 328K | 1.4M | 1.3M | |||
| Hsap UCSC200304 | 16M | 3.3M | 304M | 4.3M | 9.3M | 4.3M | 17M | 16M | |||
| Mmus UCSC200302 | 10M | 2.1M | 141M | 2.7M | 6.4M | 2.7M | 12M | 11M | |||
| Rnor UCSC200301 | 3.4M | 751K | 55M | 1M | 2.3M | 962K | 4.0M | 3.7M | |||
| U2 Both Sites | U12 Donor Site | U12 Acceptor Site | U12 Both Sites | TOTAL | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | ||||||||
| Hsap UCSC200307 | 189656 | 1529 | 34 | 2248 | 128 | 2 | 9 | 8 | 12430 | 109 | 13 | 134 | 355 | 1 | 139 | 19 | 206814 | ||||||
| Mmus UCSC200310 | 125587 | 1015 | 21 | 2407 | 88 | 0 | 7 | 9 | 9557 | 66 | 10 | 130 | 254 | 1 | 91 | 15 | 139258 | ||||||
| Rnor UCSC200306 | 38601 | 289 | 14 | 1236 | 20 | 0 | 1 | 1 | 3038 | 19 | 4 | 77 | 69 | 0 | 20 | 4 | 43393 | ||||||
| Ggal UCSC200402 | 11073 | 77 | 5 | 736 | 7 | 0 | 1 | 0 | 676 | 6 | 0 | 27 | 17 | 0 | 5 | 2 | 12632 | ||||||
| Hsap UCSC200304 | 162740 | 1254 | 28 | 2273 | 115 | 0 | 9 | 6 | 10846 | 91 | 13 | 126 | 302 | 1 | 108 | 19 | 177931 | ||||||
| Mmus UCSC200302 | 92487 | 721 | 16 | 3740 | 69 | 0 | 6 | 9 | 7027 | 46 | 5 | 192 | 196 | 1 | 67 | 9 | 104591 | ||||||
| Rnor UCSC200301 | 32378 | 253 | 13 | 1589 | 18 | 0 | 1 | 2 | 2604 | 17 | 3 | 82 | 60 | 0 | 20 | 3 | 37043 | ||||||
| Search parameters: |
donor_pattern | = | /^ATCCT[CT]/ |
|---|---|---|---|
acceptor_max_mismatch_number | = | 1 | |
acceptor_pattern | = | /TCCTT[AG]AC/ |
| U2 Both Sites | U12 Donor Site | U12 Acceptor Site | U12 Both Sites | TOTAL | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | ||||||||
| Hsap UCSC200307 | 190632 | 1536 | 34 | 2249 | 128 | 2 | 9 | 8 | 11454 | 102 | 13 | 133 | 355 | 1 | 139 | 19 | 206814 | ||||||
| Mmus UCSC200310 | 126409 | 1021 | 21 | 2408 | 89 | 0 | 7 | 9 | 8735 | 60 | 10 | 129 | 253 | 1 | 91 | 15 | 139258 | ||||||
| Rnor UCSC200306 | 38848 | 289 | 14 | 1238 | 20 | 0 | 1 | 1 | 2791 | 19 | 4 | 75 | 69 | 0 | 20 | 4 | 43393 | ||||||
| Ggal UCSC200402 | 11150 | 78 | 5 | 736 | 7 | 0 | 1 | 0 | 599 | 5 | 0 | 27 | 17 | 0 | 5 | 2 | 12632 | ||||||
| Search parameters: |
donor_pattern | = | /^ATCCT[CT]/ |
|---|---|---|---|
acceptor_max_mismatch_number | = | 2 | |
acceptor_pattern | = | /TCCTT[AG]AC/ |
| U2 Both Sites | U12 Donor Site | U12 Acceptor Site | U12 Both Sites | TOTAL | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | ||||||||
| Hsap UCSC200307 | 108118 | 973 | 21 | 1571 | 34 | 0 | 3 | 3 | 93968 | 665 | 26 | 811 | 449 | 3 | 145 | 24 | 206814 | ||||||
| Mmus UCSC200310 | 69628 | 567 | 13 | 1647 | 22 | 0 | 1 | 2 | 65516 | 514 | 18 | 890 | 320 | 1 | 97 | 22 | 139258 | ||||||
| Rnor UCSC200306 | 20943 | 168 | 9 | 855 | 4 | 0 | 0 | 0 | 20696 | 140 | 9 | 458 | 85 | 0 | 21 | 5 | 43393 | ||||||
| Ggal UCSC200402 | 6444 | 49 | 4 | 600 | 0 | 0 | 0 | 0 | 5305 | 34 | 1 | 163 | 24 | 0 | 6 | 2 | 12632 | ||||||
| Search parameters: |
donor_pattern | = | /^ATCCT[CT]/ |
|---|---|---|---|
acceptor_max_mismatch_number | = | 2 | |
acceptor_pattern | = | /TCCTT[AG]AC/ | |
| Extra constraints: |
branchpoint_distance_from_acceptor | = | [ -20 .. -5 ] |
branchpoint_sequence_matches_to | = | [ /..A.$/ || /.A..$/ ] |
| U2 Both Sites | U12 Donor Site | U12 Acceptor Site | U12 Both Sites | TOTAL | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | ||||||||
| Hsap UCSC200307 | 182013 | 1471 | 31 | 2127 | 51 | 0 | 3 | 4 | 20073 | 167 | 16 | 255 | 432 | 3 | 145 | 23 | 206814 | ||||||
| Mmus UCSC200310 | 120700 | 968 | 20 | 2316 | 32 | 0 | 1 | 2 | 14444 | 113 | 11 | 221 | 310 | 1 | 97 | 22 | 139258 | ||||||
| Rnor UCSC200306 | 37208 | 275 | 14 | 1204 | 8 | 0 | 0 | 0 | 4431 | 33 | 4 | 109 | 81 | 0 | 21 | 5 | 43393 | ||||||
| Ggal UCSC200402 | 10698 | 76 | 5 | 733 | 2 | 0 | 0 | 0 | 1051 | 7 | 0 | 30 | 22 | 0 | 6 | 2 | 12632 | ||||||
| CLASS | DONOR SITES | ACCEPTOR SITES |
|---|---|---|
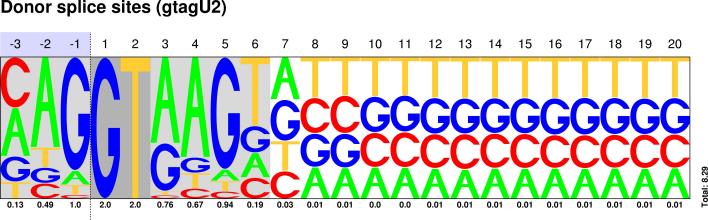
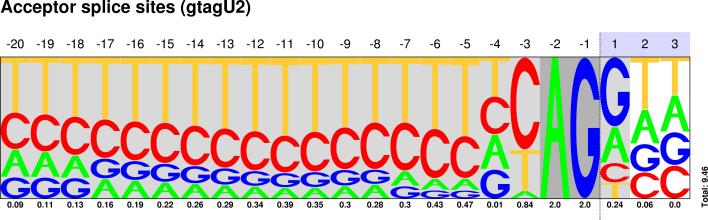
| GT-AG |
 Sequences : PWM : JPG / PNG / PS |
 Sequences : PWM : JPG / PNG / PS |
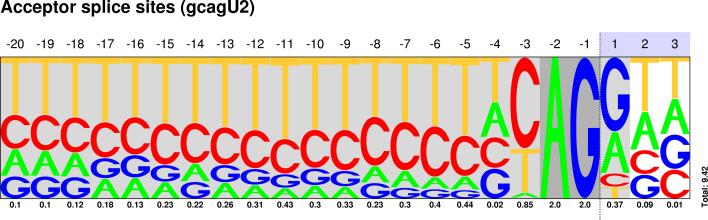
| GC-AG |
 Sequences : PWM : JPG / PNG / PS |
 Sequences : PWM : JPG / PNG / PS |
| U12 |
 Sequences : PWM : JPG / PNG / PS |
 Sequences : PWM : JPG / PNG / PS |
| BRANCH POINT |
 Sequences : PWM : JPG / PNG / PS |
|
| U2 Both Sites | U12 Donor Site | U12 Acceptor Site | U12 Both Sites | TOTAL | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | ||||||||
| Hsap UCSC200307 | 31425 | 218 | 3 | 29 | 27 | 0 | 0 | 2 | 2055 | 12 | 0 | 7 | 4 | 0 | 1 | 0 | 33783 | ||||||
| Mmus UCSC200310 | 28168 | 207 | 2 | 70 | 23 | 0 | 0 | 0 | 2231 | 14 | 1 | 9 | 2 | 0 | 0 | 0 | 30727 | ||||||
| Rnor UCSC200306 | 10019 | 64 | 4 | 23 | 5 | 0 | 0 | 1 | 835 | 9 | 0 | 5 | 0 | 0 | 0 | 0 | 10965 | ||||||
| Hsap UCSC200304 | 31626 | 220 | 3 | 28 | 27 | 0 | 0 | 2 | 2068 | 12 | 0 | 6 | 2 | 0 | 0 | 0 | 33994 | ||||||
| Mmus UCSC200302 | 28810 | 212 | 2 | 41 | 24 | 0 | 0 | 0 | 2270 | 14 | 0 | 7 | 3 | 0 | 0 | 0 | 31383 | ||||||
| Rnor UCSC200301 | 10209 | 65 | 4 | 7 | 5 | 0 | 0 | 1 | 841 | 9 | 0 | 4 | 0 | 0 | 0 | 0 | 11145 | ||||||
| U2 Both Sites | U12 Donor Site | U12 Acceptor Site | U12 Both Sites | TOTAL | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | GTAG | GCAG | ATAC | XXXX | ||||||||
| Hsap UCSC200307 | 2 | 0 | 0 | 0 | 9 | 0 | 0 | 1 | 7 | 0 | 1 | 1 | 65 | 0 | 31 | 0 | 117 | ||||||
| Mmus UCSC200310 | 1 | 0 | 0 | 0 | 2 | 0 | 2 | 0 | 7 | 0 | 2 | 1 | 71 | 0 | 27 | 1 | 114 | ||||||
| Rnor UCSC200306 | 1 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 26 | 0 | 9 | 0 | 39 | ||||||
| Hsap UCSC200304 | 0 | 0 | 0 | 0 | 10 | 0 | 0 | 1 | 7 | 0 | 1 | 1 | 67 | 0 | 31 | 0 | 118 | ||||||
| Mmus UCSC200302 | 1 | 0 | 0 | 0 | 2 | 0 | 2 | 0 | 6 | 0 | 2 | 1 | 73 | 0 | 28 | 1 | 116 | ||||||
| Rnor UCSC200301 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 27 | 0 | 9 | 0 | 39 | ||||||
| x Gg200402 | U2 Both Sites | U12 Donor Site | U12 Acceptor Site | U12 Both Sites | TOTAL | ||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Exon- erate | Genic CDS | GT AG | GC AG | AT AC | XX XX | GT AG | GC AG | AT AC | XX XX | GT AG | GC AG | AT AC | XX XX | GT AG | GC AG | AT AC | XX XX | ||||||||||
| Hs200307/Mm200310/Rn200306 | TBL | FA | GFF | 1 | 2 | 0 | 27 | 9 | 0 | 0 | 0 | 4 | 2 | 0 | 5 | 29 | 0 | 8 | 2 | 89 | |||||||
| Hs200304/Mm200302/Rn200301 | TBL | FA | GFF | 1 | 2 | 0 | 28 | 9 | 0 | 0 | 0 | 5 | 2 | 0 | 6 | 30 | 0 | 8 | 3 | 94 | |||||||
| Exonerate parameters: |
--query NNN.u12.exoncdspairs.fa | (where NNN was hsap.gp200307, mmus.gp200310, rnor.gp200306 hsap.gp200304, mmus.gp200302, or rnor.gp200301) |
|---|---|---|
--target chromfa/chrNNN.fa | (where NNN is a chicken chromosome number from this table) | |
--softmasktarget | ||
--model coding2genome | ||
--proteinsubmat blosum62 |
|
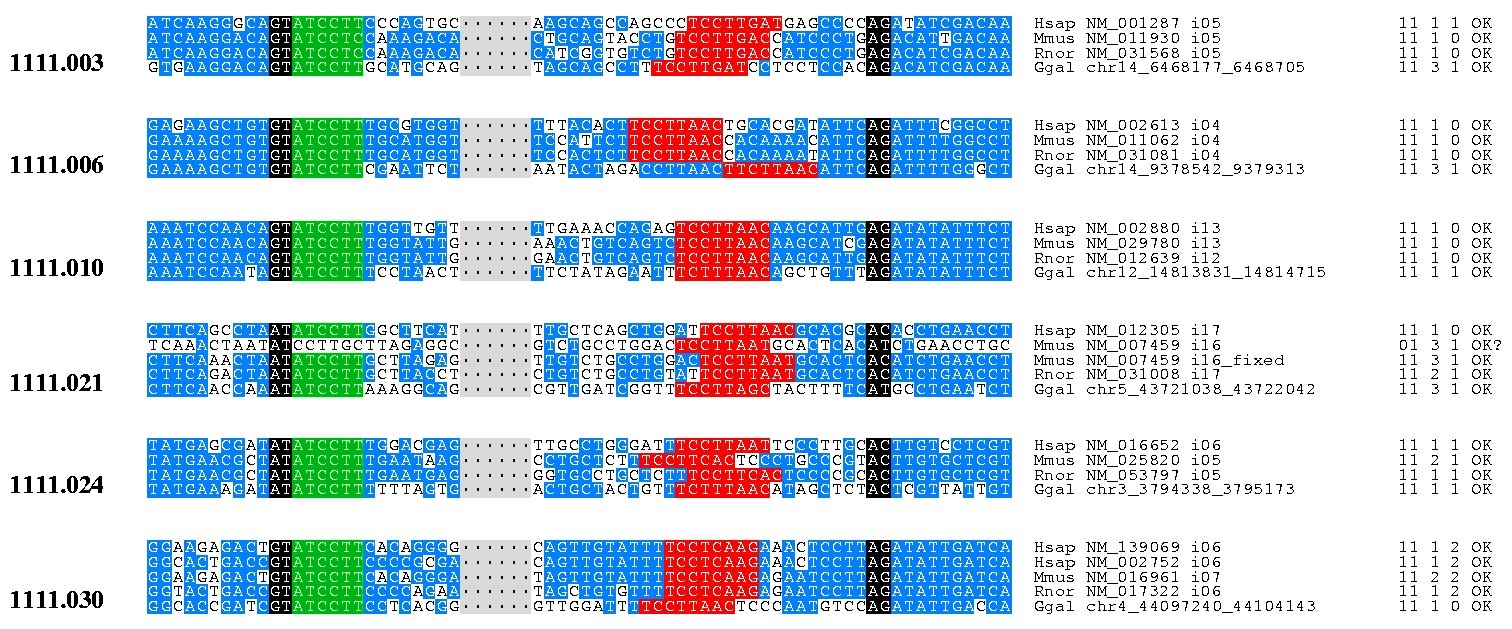
Orthologous Intron IDs Hsap/Mmus/Rnor Hsap/Mmus Hsap/Rnor Mmus/Rnor Hsap/Ggal Mmus/Ggal Rnor/Ggal Orthologous Introns IDs Hsap/Mmus/Rnor/Ggal Orthologous Splice Sites Alignments (Raw text)
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Site Sequences | |||||||
|---|---|---|---|---|---|---|---|
| Species | Data Sets | Donors (-3/GT/+4) |
Acceptors (-18/AG/+3) |
||||
| H.sapiens | seq.gz | dat.gz | dat.gz | ||||
| M.musculus | seq.gz | dat.gz | dat.gz | ||||
| R.norvegicus | seq.gz | dat.gz | dat.gz | ||||
| G.gallus | seq.gz | dat.gz | dat.gz | ||||
| D.rerio | fasta.gz | dat.gz | dat.gz | ||||
| D.melanogaster | fasta.gz | dat.gz | dat.gz | ||||
| Species | Donor Sites | Acceptor Sites |
|---|---|---|
| M.musculus R.norvegicus |
 PWM : JPG / PNG / PS |
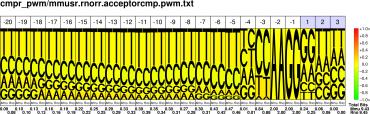 PWM : JPG / PNG / PS |
| H.sapiens M.musculus |
 PWM : JPG / PNG / PS |
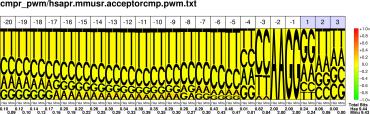 PWM : JPG / PNG / PS |
| H.sapiens R.norvegicus |
 PWM : JPG / PNG / PS |
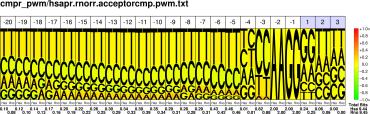 PWM : JPG / PNG / PS |
| H.sapiens G.gallus |
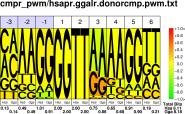 PWM : JPG / PNG / PS |
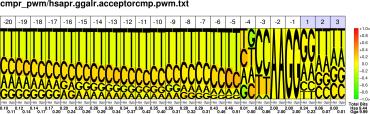 PWM : JPG / PNG / PS |
| H.sapiens D.rerio |
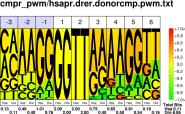 PWM : JPG / PNG / PS |
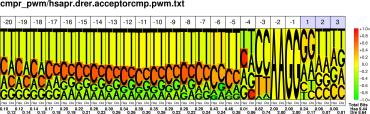 PWM : JPG / PNG / PS |
| H.sapiens D.melanogaster |
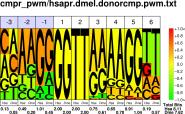 PWM : JPG / PNG / PS |
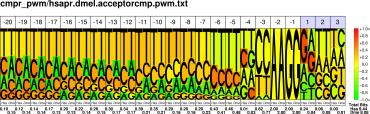 PWM : JPG / PNG / PS |
To perform the following analyses we started from a set of human, mouse, rat and chicken, reliable 1:1:1:1 orthologs, kinly provided by Peer Bork and Ivica Letunic as part of the International Chicken Genome Sequencing Consortium (ICGSC) collaborations. From that set we produced the file linked below, containing the 1:1:1:1 orthologous introns for which the donor and acceptor sites used in the conservation analysis were retrieved.
orthointrons_sites.hmrg.tbl contains 6524 orthologous introns.
| Based on | All Exons | All Introns | All CDSs | Splice Sites | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ensgenes.txt | SEQ(fasta) | SEQ(content) | SEQ(fasta) | SEQ(content) | SEQ(fasta) | SEQ(content) | EXONIC | INTRONIC | |||
| Hsap UCSC200307 | 22M | 5.0M | 507M | 6.2M | 24M | 6.2M | 22M | 20M | |||
| Mmus UCSC200310 | 20M | 4.2M | 293M | 5.1M | 21M | 5.3M | 19M | 18M | |||
| Rnor UCSC200306 | 13M | 3.7M | 234M | 4.4M | 14M | 4.1M | 17M | 15M | |||
| Ggal UCSC200402 | 13M | 4.1M | 180M | 4.7M | 14M | 4.6M | 18M | 16M | |||
| Donor Sites | Acceptor Sites | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Species | Orthologous Pairs |
Identity Summary |
Random Pairs |
Identity Summary |
Orthologous Pairs |
Identity Summary |
Random Pairs |
Identity Summary |
|||||||
| M.musculus | / | R.norvegicus |
TBL | SET | TBL | SET | TBL | SET | TBL | SET | |||||
| H.sapiens | / | M.musculus |
TBL | SET | TBL | SET | TBL | SET | TBL | SET | |||||
| H.sapiens | / | R.norvegicus |
TBL | SET | TBL | SET | TBL | SET | TBL | SET | |||||
| H.sapiens | / | G.gallus |
TBL | SET | TBL | SET | TBL | SET | TBL | SET | |||||
| G.gallus | / | M.musculus | TBL | SET | TBL | SET | TBL | SET | TBL | SET | |||||
| G.gallus | / | R.norvegicus |
TBL | SET | TBL | SET | TBL | SET | TBL | SET | |||||
| Human/mouse/rat/chicken orthologous introns file: TBL | |||||||||||||||