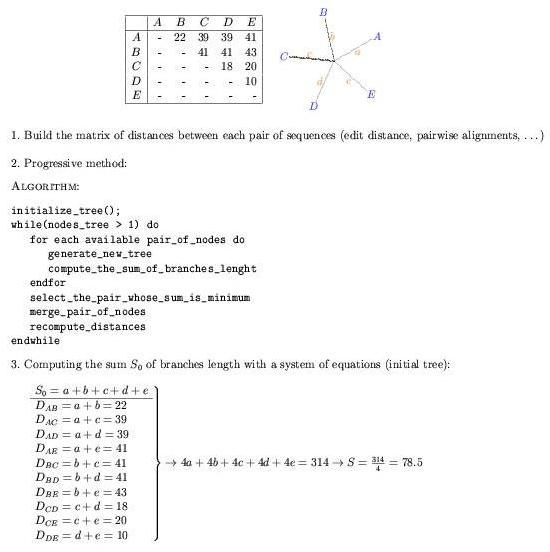
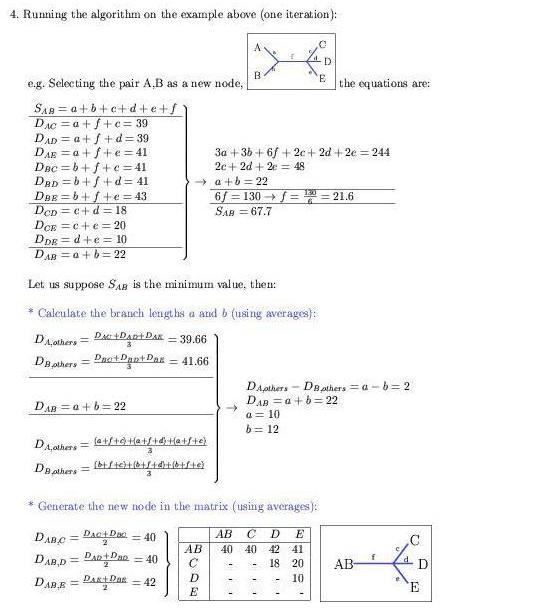
Distance based methods: Minimize the number of global changes
between each pair in a group of sequences previously aligned.
Least squares approach:
From a matrix of distances D between species, find a phylogenetic tree
that predicts a set of distances d so that the following expression is
minimized:
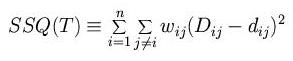
Neighbour-joining:
- Join the clusters that are close to another and also apart from the rest
- Heuristic: Minimize the sum of the branch lengths in the tree
- Additivity of evolutionary distances

